In our March blog post, we discussed why population sizes are important for conserving genetic biodiversity, why we use the effective population size Ne (rather than just a count of individuals), and what exactly that is. In today’s post, we’ll get into how to obtain Ne estimates in practice!
As a quick reminder: The Ne 500 indicator was developed to assess whether populations are large enough to maintain genetic variation in the long term. This indicator needs to be reported to the Convention on Biological Diversity by all member states from this year onwards. It is defined as the proportion of populations within species with Ne > 500. Ne, estimated for as many populations of a species as possible, is therefore the central “ingredient” of this indicator.
The threshold of 500 is the result of extensive research showing that a population of this effective size can largely maintain genetic diversity over time (Jamieson & Allendorf 2012). Populations with Ne over 500 do still lose genetic diversity over time – but this loss is slow and is partly compensated for by mutation, which generates new variation.
How do I get estimates of Ne?
You have quite a range of options for obtaining Ne estimates (arrows in different colours in the figure below): You can use different types and sources of data (light turquoise fields), and you can either obtain Ne directly or determine it from an estimate of the census size, Nc.
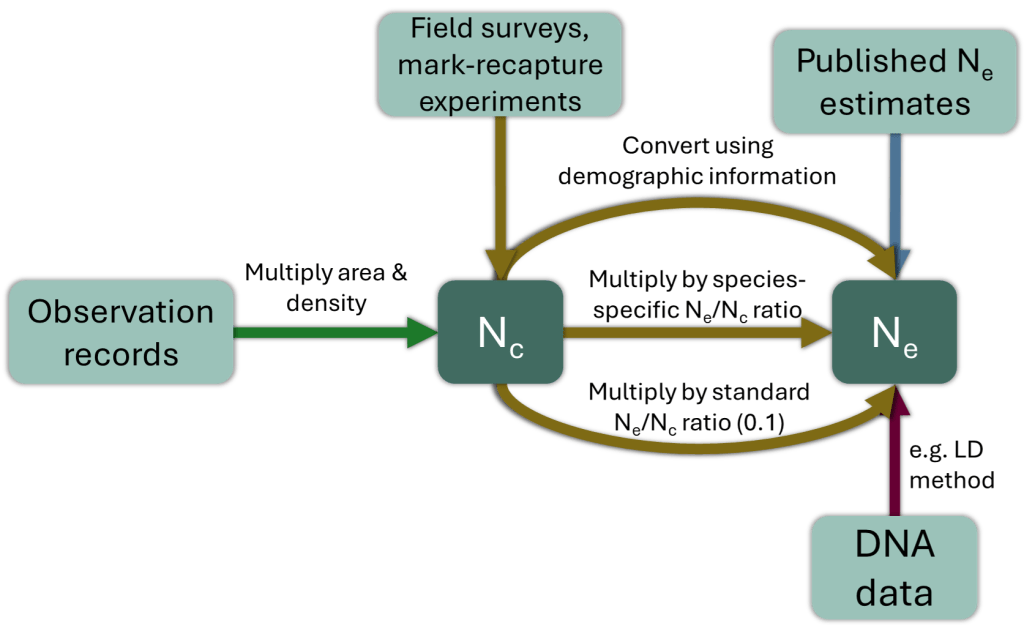
The (relatively) easy way:
Ne estimates can sometimes be found in the scientific literature – you can just gather those values and plug them into your indicator calculation. Important caveat: Some published estimates cover genetic patterns over hundreds or thousands of generations into the past. Such estimates are not useful in assessing genetic erosion today. What matters in the conservation context is the contemporary Ne. Suitable estimates are those based on the Linkage Disequilibrium or Sibship methods, for example. Additional acceptable methods are listed here.
The DNA way:
If DNA data are available, e.g., microsatellite or SNP (single nucleotide polymorphism) data, Ne can be estimated directly from the extent of genetic changes over time or from the associations between variants at different genes (Linkage Disequilibrium). A range of software programs exists, and GINAMO is working on a pipeline to automate the estimation process as much as possible. We recommend the Linkage Disequilibrium method because it does not require sampling over time and is relatively reliable.
The indirect way:
It is possible to first obtain census population size (Nc) estimates – e.g., from actual individual counts or mark-recapture studies – and then convert them into Ne:
- If the Ne/Nc ratio for the species is known – for example from the scientific literature – Ne can be estimated by using the formula Ne = Nc * (Ne/Nc). More information on the Ne/Nc ratio can be found in our March blog post.
- If the Ne/Nc ratio for the species is unknown, the “default” value of 0.1 can be used. For most species, this is roughly correct and tends to be conservative (Hoban et al. 2020). It is recommended to try several Ne/Nc ratios if you are unsure about the most likely range for your species; this is being explored in a GINAMO subproject and will be the topic of a future post.
- If factors such as variation in offspring number between families have been quantified, they can be used in equations from population genetics theory to calculate Ne (Waples 2025). One example of this approach is a study on a well-monitored Norwegian reindeer population.
The more indirect way:
When neither DNA data nor Nc data are available, it may still be possible to roughly estimate Ne – or at least to check whether Ne is likely greater than 500 – based on distribution data. This approach allows for an assessment of populations that haven’t been studied in detail. First, obtain the area occupied by the population, which can often be extracted from species observation records, e.g., in GBIF or iNaturalist. Multiply it with the typical density of the species (number of individuals per area, often available from the literature or species experts). This way, you get a rough estimate of Nc, which you can use to estimate Ne in one of the ways described above. This value may not be accurate, but it can often be enough to assess whether the true Ne is likely much smaller or larger than 500.

In reindeer, the effective population size has been estimated from demographic data.
What do I then do with my Ne estimates?
Once Ne is available for the different populations of a species, the species-level Ne 500 indicator can easily be calculated – it is just the number of populations with Ne > 500 divided by the total number of populations assessed. This indicator can be reported to the CBD at the species level; in addition, it contributes to the country-level indicator that combines all species.
Like all biodiversity indicators, Ne 500 is a simplified measure – it doesn’t tell us everything there is to know about a species’ genetic status and risk. Some populations with Ne greater than 500 might be declining rapidly (for non-genetic reasons) and could already have lost relevant adaptive variants, while in other populations even Ne smaller than 500 might not be a cause for concern (e.g., because they have persisted naturally at this size for long periods). For these reasons, it is important to store and keep track of the estimated Ne values (and, if available, Nc values) over time, not just the indicator values. Ne alone can also not capture all genetic processes (Allendorf et al. 2024, Mergeay 2024), and the association between Ne and genetic diversity isn’t perfect. A population can be large now but still have little genetic variation if it recently went through a period when it was much smaller. In such cases, Ne reflects whether the population is large enough to maintain its current genetic diversity, but not the extent of that diversity.
The Ne 500 indicator is a valuable tool for tracking trends and identifying populations at risk. To reach its full potential, the underlying data (the actual Ne estimates, as well as other measures of genetic diversity and structure) should be stored and analysed in the context of population history and ecology.
More information about estimating Ne can be found in the guideline materials produced by the Coalition for Conservation Genetics.

